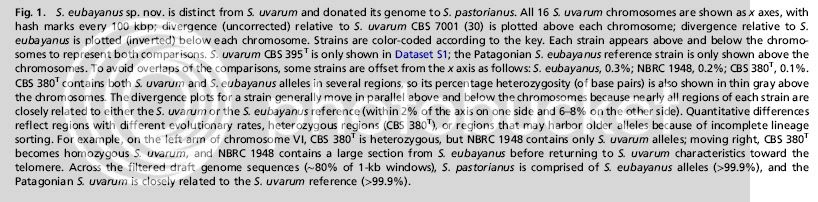
Good to know a scientist with research paper access.
 Microbe domestication and the identification of the
Microbe domestication and the identification of the
wild genetic stock of lager-brewing yeast
Diego Libkinda,1, Chris Todd Hittingerb,c,1,2, Elisabete Valériod, Carla Gonçalvesd, Jim Doverb,c, Mark Johnstonb,c,
Paula Gonçalvesd, and José Paulo Sampaiod,3
aLaboratorio de Microbiología Aplicada y Biotecnología, Instituto de Investigaciones en Biodiversidad y Medio-ambiente, Consejo Nacional de Investigaciones
Científicas y Técnicas (CONICET)-Universidad Nacional del Comahue, 8400 Bariloche, Argentina; bDepartment of Biochemistry and Molecular Genetics,
University of Colorado School of Medicine, Aurora, CO 80045; cDepartment of Genetics, Center for Genome Sciences, Washington University in St. Louis
School of Medicine, St. Louis, MO 63108; and dCentro de Recursos Microbiológicos, Departamento de Ciências da Vida, Faculdade de Ciências e Tecnologia,
Universidade Nova de Lisboa, 2829-516 Caparica, Portugal
Edited by John Doebley, University of Wisconsin, Madison, WI, and approved July 20, 2011 (received for review April 5, 2011)
Domestication of plants and animals promoted humanitys transition
from nomadic to sedentary lifestyles, demographic expansion,
and the emergence of civilizations. In contrast to the well-documented
successes of crop and livestock breeding, processes of microbe
domestication remain obscure, despite the importance of
microbes to the production of food, beverages, and biofuels. Lagerbeer,
first brewed in the 15th century, employs an allotetraploid
hybrid yeast, Saccharomyces pastorianus (syn. Saccharomyces
carlsbergensis), a domesticated species created by the fusion of
a Saccharomyces cerevisiae ale-yeast with an unknown cryotolerant
Saccharomyces species. We report the isolation of that species
and designate it Saccharomyces eubayanus sp. nov. because of its
resemblance to Saccharomyces bayanus (a complex hybrid of S.
eubayanus, Saccharomyces uvarum, and S. cerevisiae found only
in the brewing environment). Individuals from populations of S.
eubayanus and its sister species, S. uvarum, exist in apparent sympatry
in Nothofagus (Southern beech) forests in Patagonia, but are
isolated genetically through intrinsic postzygotic barriers, and ecologically
through host-preference. The draft genome sequence of
S. eubayanus is 99.5% identical to the non-S. cerevisiae portion of
the S. pastorianus genome sequence and suggests specific changes
in sugar and sulfite metabolism that were crucial for domestication
in the lager-brewing environment. This study shows that combining
microbial ecology with comparative genomics facilitates
the discovery and preservation of wild genetic stocks of domesticated
microbes to trace their history, identify genetic changes, and
suggest paths to further industrial improvement.
beer yeast | next-generation sequencing | yeast ecology | yeast taxonomy
The beginning of agriculture and the domestication of plants
and animals are among the most decisive events in human
history because they triggered the rise of civilizations and the
attendant demographic, technological, and cultural developments
(1). The domestication of barley in the Fertile Crescent
(2) led to the emergence of the forebear of modern beer in
Sumeria 6,000 y ago (3). Beer and other alcoholic beverages
may have played a pivotal role in cementing human societies
through the social act and rituals of drinking (4) and by providing
a source of nutrition, medicine, and uncontaminated
water (5). Since the emergence of fermented beverages roughly
matches the domestication of plants and animals, it is likely that
some yeast lineages with favored traits were also unwittingly
domesticated.
In Europe, brewing gradually evolved during the Middle Ages
to produce ale-type beer, a process conducted by Saccharomyces
cerevisiae, the same species involved in producing wine and
leavened bread. Lager-brewing arose in 15th century Bavaria,
gained broad acceptance by the late 19th century (6), and has
since become the most popular technique for producing alcoholic
beverages, with over 250 billion dollars of global sales in
2008 (7). Unlike most ales and wines, lagers require slow, lowtemperature
fermentations that are carried out by cryotolerant
Saccharomyces pastorianus (syn. Saccharomyces carlsbergensis)
strains (8); two other cryotolerant Saccharomyces spp. have been
associated with beer as contaminants (Saccharomyces bayanus)
and with cider or wine fermented at low temperatures (Saccharomyces
uvarum) (9). S. pastorianus has never been isolated from
the wild, depends on humans for its propagation, and appears to
be an allotetraploid hybrid species of S. cerevisiae and an unidentified
species (10, 11). Several hypotheses have been advanced
for the source of the non-S. cerevisiae genome present in
S. pastorianus, including the taxonomically and genetically complex
species S. bayanus (1214) and an unknown lager lineage
distinct both from S. bayanus and S. uvarum (11, 15). Identifying
the wild genetic stock of the cryotolerant subgenome of S. pastorianus
is necessary for resolving the taxonomy and systematics
of this important species complex, and for understanding the key
events that led to the domestication of lager yeast.
In contrast to extensive investigation into domestication of
crops and livestock (2, 1619), studies of domestication of
eukaryotic microbes have been limited (2024), perhaps because
of the inability to conduct direct field studies. Identifying the
genetic basis of traits under selection during domestication may
clarify the emergence of new traits and show the way toward
further improvement. Because domesticated lineages derive
from a subset of the original populations, a genetic bottleneck is
likely to have caused the disappearance of some alleles (17),
especially in microbes, which are often propagated clonally. In an
age of accelerated habitat destruction and diminishing biodiversity,
discovery of wild genetic stocks of domesticated
microbes will facilitate preservation of their genetic resources for
strain improvement.
Results and Discussion
Discovery of Wild Populations of Cryotolerant Saccharomyces. Saccharomyces
spp. are associated with oak trees (Fagaceae) in the
Northern Hemisphere (25, 26). Because species of the genus
Nothofagus (Southern beeches, also members of the Fagales)
occupy the oak niche in temperate regions of the Southern
Hemisphere (27), our survey in Northwestern Patagonia for Saccharomyces focused on woodlands containing populations of
Nothofagus antarctica, Nothofagus dombeyi, and Nothofagus pumilio,
within and near Lanin and Nahuel Huapi National Parks
(Argentina) (Fig. S1). We also surveyed stromata of Cyttaria hariotii
(an obligate ascomycete parasite of Nothofagus spp.) because
these fruiting structures are rich in simple sugars and provide
a favorable yeast habitat (28).Atotal of 133 samples of Nothofagus
bark, soil from underneath the trees, and Cyttaria stromata, collected
from 2006 to 2008, yielded 123 isolates of cryotolerant
Saccharomyces and two isolates of S. cerevisiae (Table S1).
For a preliminary identification of the cryotolerant Saccharomyces
isolates, we determined the DNA sequence of individual
genes, performed PCR-fingerprinting, and examined restriction
fragment length polymorphisms (RFLPs), using S. bayanus CBS
380T, S. uvarum CBS 395T, and S. uvarum CBS 7001 (referred to
as S. bayanus in genomics literature) as references (Fig. S2). The
isolates discretely fall into two groups: group A appears related
to S. bayanus (78 isolates); group B is closely related to S. uvarum
(45 isolates). The almost complete occupancy of the Nothofagus
niche by cryotolerant species contrasts with our ongoing survey
of Saccharomyces biogeography in the Northern Hemisphere oak
niche (North America, Mediterranean, Central Europe, and
Japan), where we have isolated ∼240 Saccharomyces strains from
more than 500 oak samples and observed that sympatric species
tend to have different growth temperature preferences. For
example, S. cerevisiae (thermotolerant) and Saccharomyces
kudriavzevii (cryotolerant) co-occur in Mediterranean regions,
but Saccharomyces paradoxus (thermotolerant) and S. uvarum
(cryotolerant) co-occur in temperate Europe and North
America (26). Therefore, the detection of a pair of cryotolerant
species in Patagonia and the near absence of thermotolerant
species suggest the Patagonian ecosystem supporting Saccharomyces
spp. may be unusual. One potential explanation is that
the relatively low annual average temperatures in our Patagonian
isolation sites [6/8 °C with mean low temperatures of −1/
−2 °C and mean high temperatures of 22/23 °C (29)] may favor
cryotolerant over thermotolerant species.